Mechanisms of GPCR Signaling |
|
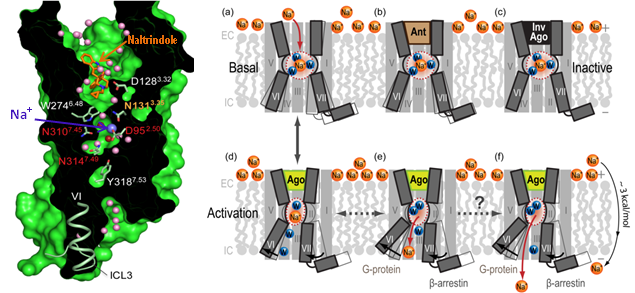
Recent breakthroughs in GPCR structural biology have led to an exponential growth in the number of GPCR crystal structures. Our studies as a part of the GPCR Network project involve comparative and conformational modeling of the receptors to answer important questions about functional mechanisms and pharmacology of GPCRs. Examples of such structure-based computational studies include: (i) deciphering molecular basis of subtype selectivity for 2nd-generation antihistamine drugs, (ii) analysis and modeling of activation-related conformational changes in Adenosine A2A receptor (AA2R), (iii) prediction of ligand binding, subtype and functional selectivity for opioid (KOR, NOP, DOR), and serotonin receptors 5HT1B and 5HT2B. Computational studies have been also crucial in predicting effects of mutations and designing new experiments to explore biological and pharmacological hypotheses using the determined structures. Most recently, computational tools have also have been applied to deciphering the functional role of a highly conserved allosteric site for binding of sodium and water cluster in the middle of the 7-transmembrane (7TM) domain of class A GPCRs. After initial discovery of the sodium site in Adenosine A2A receptor (AA2R), we predicted the presence of the conserved sodium in a majority of Class A GPCRs and analyzed the dynamics of the sodium pocket in different conformational states of receptors. Since then, this common Na+ site was confirmed in several highly diverse GPCRs, including opioid receptors. Moreover, comprehensive analysis of conformational, functional and evolutionary aspects of this conserved allosteric pocket points to a key role of this sodium cluster as a co-factor in class A GPCR signaling ( see Fig. 2 and our TiBS review).
|